Image


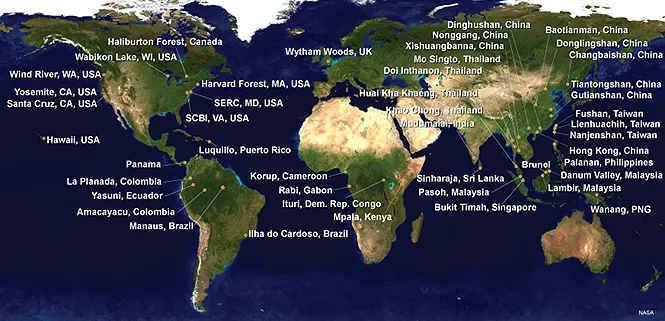
The Smithsonian National Museum of Natural History Department of Botany Plant DNA Barcode Group is collaborating with the Center for Tropical Forest Science (CTFS)/ForestGEO on an initiative to generate DNA barcodes for the trees found in the forest dynamic plots. The project will eventually generate DNA barcodes for the over 8,500 species of trees recorded from the plots. The main goals are to provide genetic identifiers for each species, test species boundaries, assist in identifying new species, provide a tool for ecological forensic applications, and generate testable phylogenetic hypotheses for the tree communities. The Smithsonian PIs are seeking collaborations with local plot PIs to initiate barcoding activities at plots around the world. This website provides information on the basic DNA barcoding protocol for forest dynamic plots. In order to standardize the collection processes, the following protocol should be used by all participants.
1) Specimen Data
An Excel file will be sent to participating CTFS sites. The specimen data entered into the spreadsheet is necessary before the Plant DNA Barcoding Group can begin the DNA extraction process.
2) Dried tissue samples in extraction blocks
Silica gel dried tissue samples (4 samples per species) should be placed in extraction blocks. Instructions will be sent on the order of loading quadruplicate samples. We can send the extraction blocks to collaborating CTFS sites, on request.
3) Duplicate tissue samples of all tissue to be extracted
Duplicate/backup silica gel dried tissue samples of all plant samples should be placed in acid free coin envelopes. We can send coin envelopes to collaborating CTFS sites, on request.
4) One duplicate pressed specimen with voucher label
The duplicate pressed specimen will be accessioned into the collections of the US Herbarium in Washington, DC, USA. It will be mounted by the US Herbarium on an herbarium sheet, assigned a unique sheet number and a unique UPC barcode number that can be used for publication purposes.